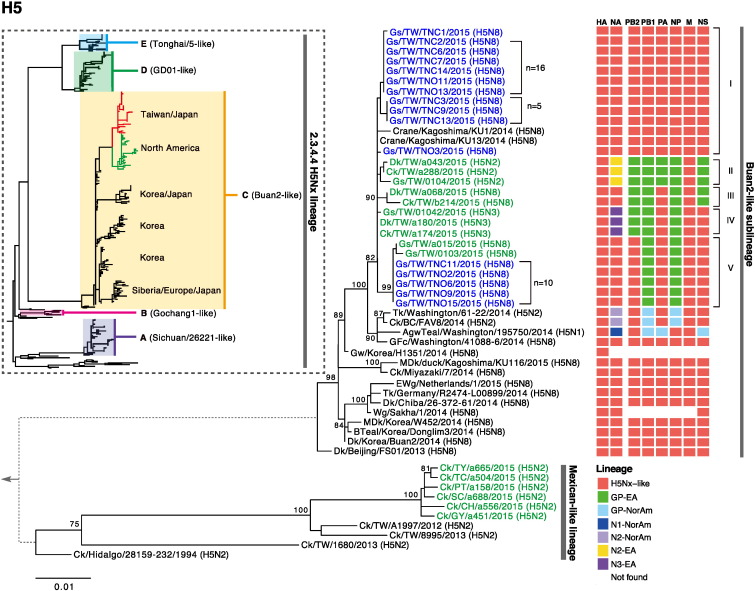
ggtree: visualization and annotation of phylogenetic trees

The ggtree package extending the ggplot2 package. It based on grammar of graphics and takes all the good parts of ggplot2. ggtree is designed for not only viewing phylogenetic tree but also displaying annotation data on the tree.
ggtree is released within the Bioconductor project and the source code is hosted on GitHub.
Citation
Please cite the following article when using ggtree:
G Yu, DK Smith, H Zhu, Y Guan, TTY Lam*. ggtree: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods in Ecology and Evolution. 2017, 8(1):28-36.
Installation
Install ggtree is easy, follow the guide on the Bioconductor page:
## try http:// if https:// URLs are not supported
source("https://bioconductor.org/biocLite.R")
## biocLite("BiocUpgrade") ## you may need this
biocLite("ggtree")If you have problems when installing some of the dependent packages, please refer to the ggtree-installation wiki page.
Overview
Getting tree into R
- tree parsers: bring evolution evidences to be used/analyzed in
R merge_tree: allows evolution evidences to be merged and comparedfortifymethods: convert tree objects into tidy data frame
Tree visualization & annotation
- parsing tree as a collection of nodes allows grammar of graphics to be supported
geom_tree: extendsggplot2to support tree structure- several layers and functions for tree annotation
- supports annotating phylogenetic trees with user’s own data
Tree manipulation
- helper functions for tree manipulation, make it possible to explore the tree visually
Find out details and examples on Documentation.
Projects that depend on ggtree
CRAN packages
- genBaRcode: Analysis and Visualization Tools for Genetic Barcode Data
- harrietr: Wrangle Phylogenetic Distance Matrices and Other Utilities
Other applications
- BreadCrumbs: Collection of scripts for metagenomics analysis
- DegeneratePrimerTools: Utilities for Creating and Validating Degenerate primers
- phyloscan: scan phylogenies created along a genome for patterns
- microbiomeViz: Visualize microbiome data
- ggnetworx: Phylogenetic networks using ggplot2 and ggtree
Feedback
- Please make sure you have followed the important guide before posting any issue/question
- For bugs or feature requests, please post to github issue
- For user questions, please post to google group
- We are also following every post tagged with ggtree on Bioconductor support site and Biostars
- Join the group chat on and